- About
-
Solutions
-
Services
- Biosciences
- Chemistry
- Integrated Drug Discovery
- Computer Aided Drug Design
- Hit Identification
- DMPK Services
- Target Classes and Modalities
- Therapeutic Areas
-
A-Z
- A
- B
- C
- D
- E
- F
- G
- H
- I
- K
- L
- M
- N
- O
- P
- R
- S
- T
- V
- X
-
Services
- Library
- News & Events
- Careers
Molecular Glues
Small single-unit protein-protein binders
Molecular glues are a class of compounds which either stabilise the interaction between two proteins or induce new protein-protein interactions (PPI). Domainex has experience working with molecular glues and has established a platform of biochemical and biophysical assays, for hit finding and detailed characterisation.
Molecular glues work by enhancing the affinity between the two proteins and thus have enabled once deemed “undruggable” proteins to be targeted. Molecular glues achieve this by altering the surface of the bound protein in a manner that complements the surface of the binding partner, thus ‘gluing’ the two targets together. This interaction can either stabilise endogenous PPIs or induce non-native ones. There are several molecular glues approved by the FDA.
Where one of the PPI binders is an E3 ligase, molecular glues promote targeted protein degradation by recruiting ubiquitin ligases in a similar way to proteolysis targeting chimeras (PROTACs®). However, whilst molecular glues bear similarities to PROTACs®, there are some key differences. For example, molecular glues are comprised of one unit, which forms highly co-operative binding between the two proteins, whereas PROTACs® are bivalent heterobifunctional molecules consisting of two ligands (one which binds to the target protein and the other to the E3 ubiquitin ligase) joined by a linker. Therefore, molecular glues are typically significantly smaller than PROTACs®, meaning they are well suited to the development of compounds with physio-chemical properties compatible with oral delivery and/or CNS targets . An additional added benefit of their structural simplicity lies in their more straight forward synthesis and purification in comparison to PROTACs®.
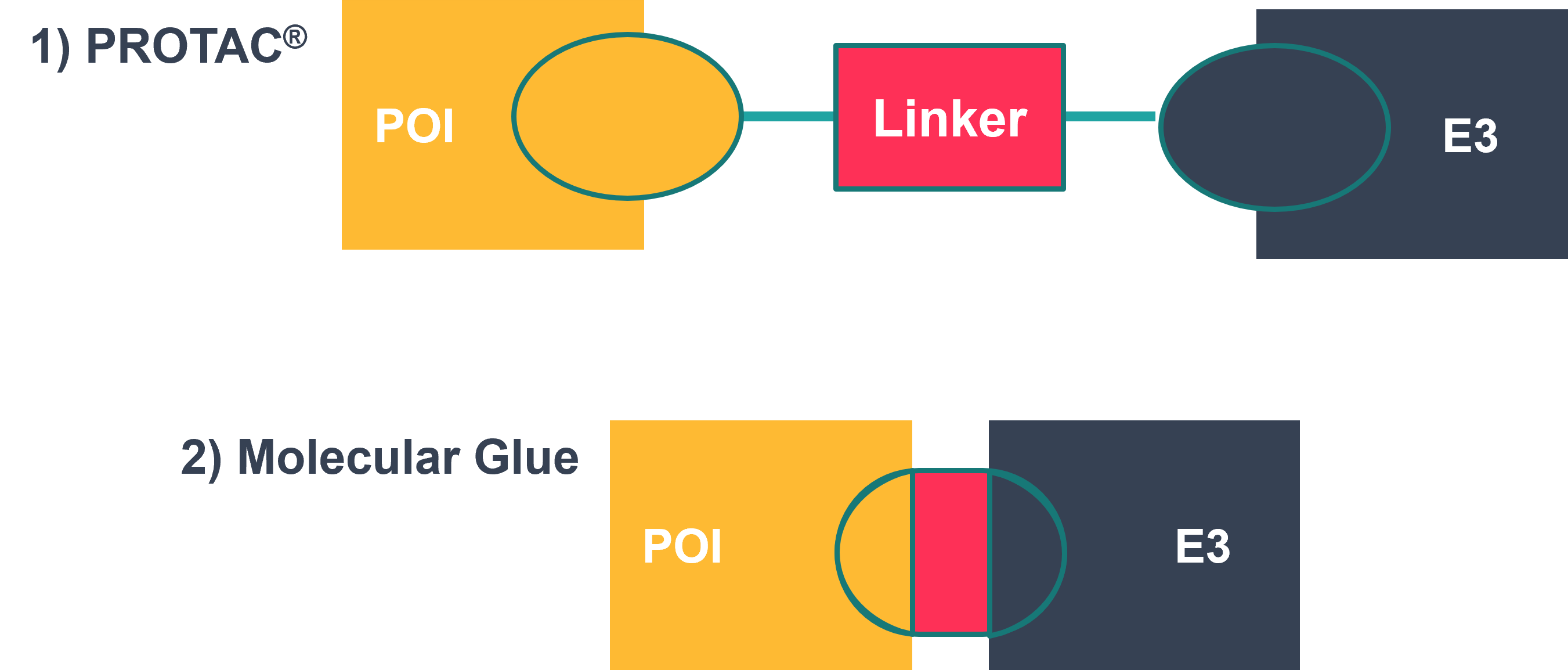
Figure 1: Schematic of protein degradation by a 1) PROTAC® and 2) molecular glue
Molecular Glues at Domainex
At Domainex, we are experienced in hit finding, design, synthesis and profiling of molecular glues and we always tailor our approach to the specific needs of your project.
Hit finding for novel molecular glues is challenging because there is a requirement for molecular glues is to enhance the formation of a ternary complex, meaning compounds are required to enhance cooperativity rather than binding to a single target. Therefore, fragment screening is an attractive hit identification approach and our FragmentBuilder screening platform is ideal for this. Typically, we use Spectral Shift or AlphaScreen to rapidly screen our in-house fragment library. Both these techniques are highly sensitive and can be deployed to screen 10,000s – 100,000s of compounds in a co-operativity assay format.
A)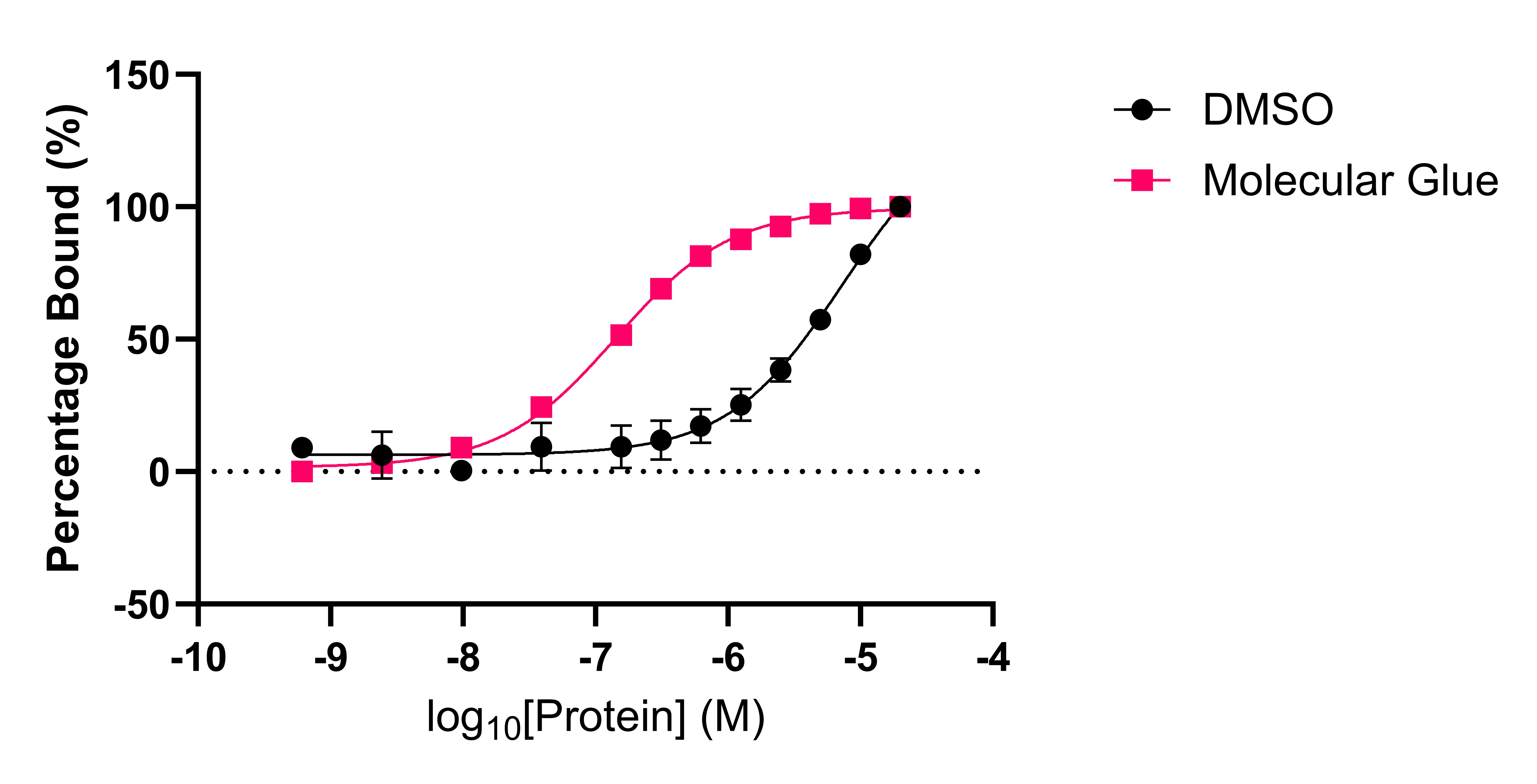 B)
B)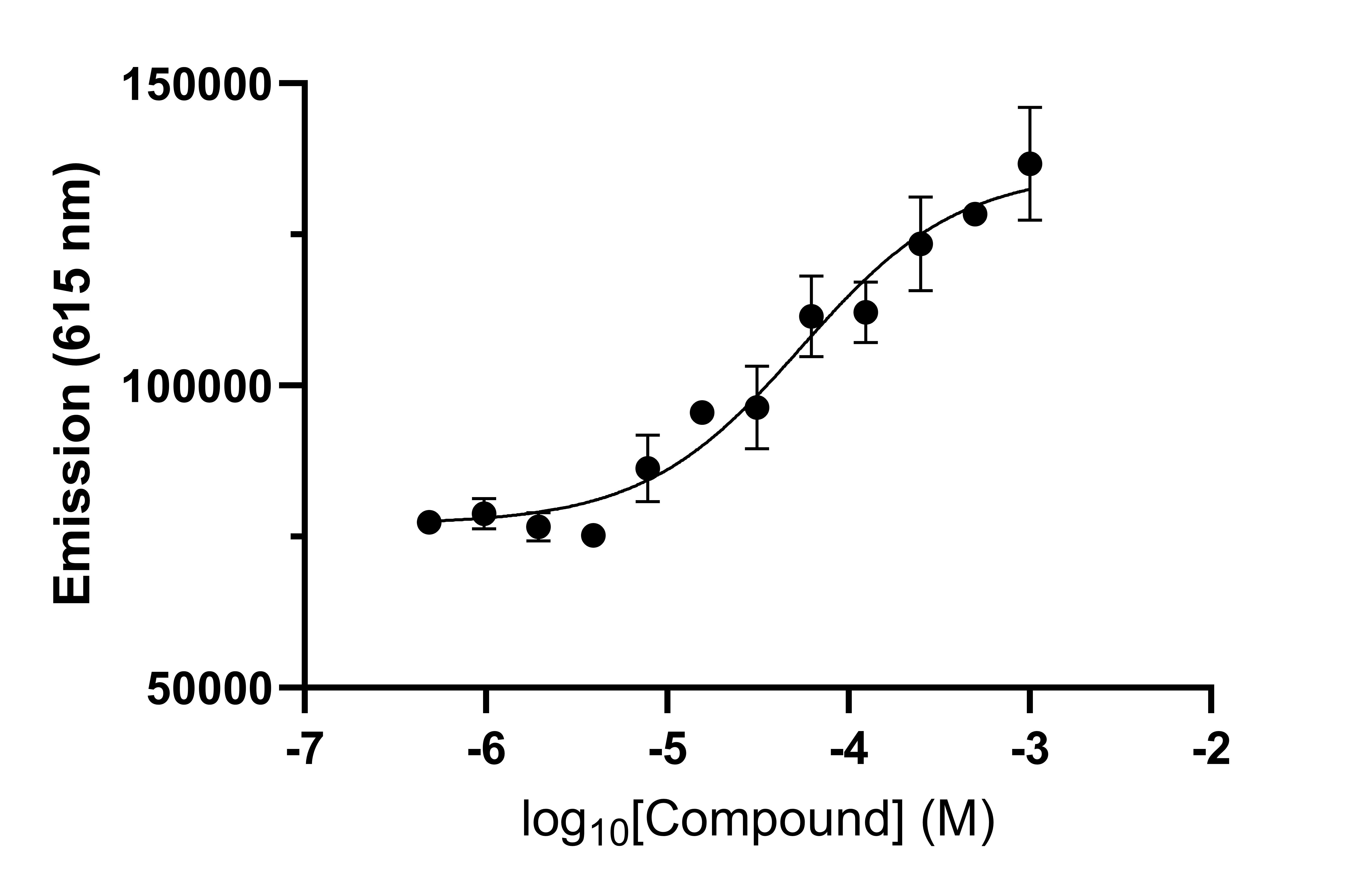
Figure 2: (A) An example of a molecular glue enhancing a weak PPI interaction via Spectral Shift (NanoTemper). The DMSO control shows a weak interaction between two proteins (black) which is enhanced approximately 50-fold (co-operativity factor a = 51.4) by a molecular glue compound (red). (B) Determination of the EC50 of a molecular glue to form a ternary complex in an AlphaScreen assay.
Hits identified from these screens can be profiled and characterised in more detail. At Domainex, we deploy our breadth of biophysical expertise and high-quality instrumentation to lower throughput orthogonal assays, such as Grating Coupled Interferometry (GCI)/Surface Plasmon Resonance (SPR) to assess binding kinetics, and mass photometry to demonstrate ternary complex formation without any labelling. X-ray crystallography or cryo-EM can also be employed to gain a structural understanding of the ternary complex and to aid drug design, while our internal protein scientists are experienced in the production of a variety of E3 ligases and we have in-house VHL and Cereblon available for immediate use.
A) 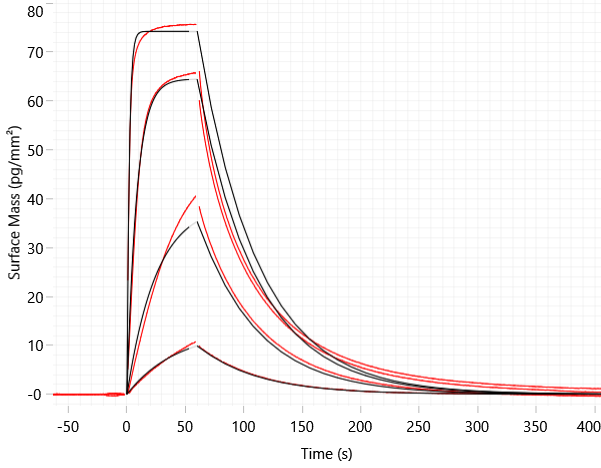 B)
B) 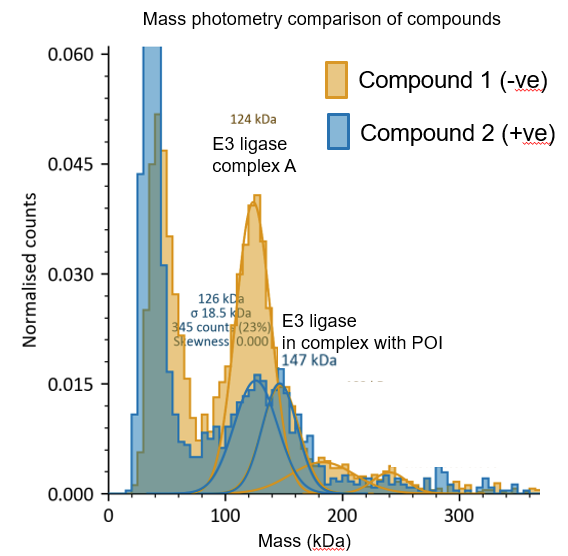
C)
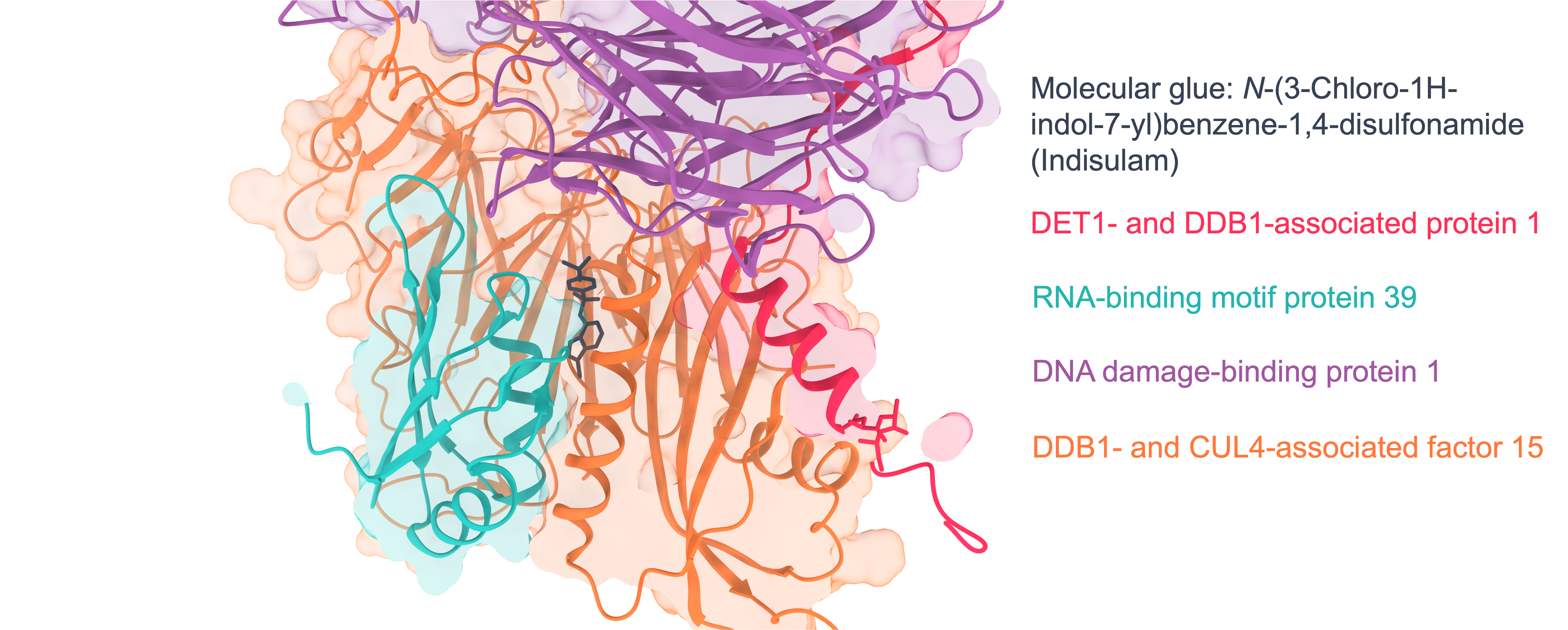
Figure 3: Molecular glue cascade; Following hit finding, orthogonal assays such as (A) GCI and (B) mass photometry can be used to confirm ternary complex formation. Structural biology can be used to identify where the molecular glue interacts via Cryo-EM or (C) X-ray crystallography (example literature image shown; PDB 6UD71).
Domainex also has a wealth of experience in running cellular degrader assays using Promega HiBiT tagged lines and automated Western Blotting for native degradation. With the appropriate counter-screens HiBiT assays can also be used for the identification of glue-degraders with proteomic techniques such as BioID or TurboID used to identify the degrading E3 ligase after hit finding.2
References
Dirkensen E. Bussiere et al., Nature Chemical Biology, 2020, 16, 15-23. https://doi.org/10.1038/s41589-019-0411-6
Holdgate GA, et al. Screening for molecular glues - Challenges and opportunities. SLAS Discov. 2024 Mar;29(2):100136. https://doi.org/10.1016/j.slasd.2023.12.008
PROTAC® is a registered trademark of Arvinas Operations, Inc., and is used under license.
TernaryTx partnered with Domainex to advance two of our molecular glue discovery programs. Specifically, they carried out a campaign utilising their cell-based biology capabilities and protein degradation expertise to identify hits against novel targets identified by the TernaryTx platform.
Our project team at Domainex impressed us with their scientific rigour, excellent communication, and collaborative approach to problem-solving. The team's flexibility with tight deadlines and changing priorities has been invaluable, helping us successfully validate hits against a classically 'undruggable' target and accelerate our drug discovery efforts.
Our project team at Domainex impressed us with their scientific rigour, excellent communication, and collaborative approach to problem-solving. The team's flexibility with tight deadlines and changing priorities has been invaluable, helping us successfully validate hits against a classically 'undruggable' target and accelerate our drug discovery efforts.
Dr Chris Tame - CEO and Dr Adam Yip - project lead, TernaryTx
Start your next project with Domainex
Contact one of our experts today